This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
Homology
What is Homology?
Homology is a similarity between different species across many different features such as structure or physiology (1). Characteristics or structures are considered to be homologous if they have evolved from the same characteristic or structure in their common ancestor (2). In order to conduct research using model organisms to find information about the human condition, it is important to understand how similar the structures, physiology, and genetics are between organisms (2). Indeed, the foundation of genetic homology is especially important when using model organisms to study human conditions as small changes to the nucleotide sequence could bring about big changes in the expression of the gene, leading to gaps between the model and human.
Homology of NDN
Homology is a similarity between different species across many different features such as structure or physiology (1). Characteristics or structures are considered to be homologous if they have evolved from the same characteristic or structure in their common ancestor (2). In order to conduct research using model organisms to find information about the human condition, it is important to understand how similar the structures, physiology, and genetics are between organisms (2). Indeed, the foundation of genetic homology is especially important when using model organisms to study human conditions as small changes to the nucleotide sequence could bring about big changes in the expression of the gene, leading to gaps between the model and human.
Homology of NDN
Blast Results:
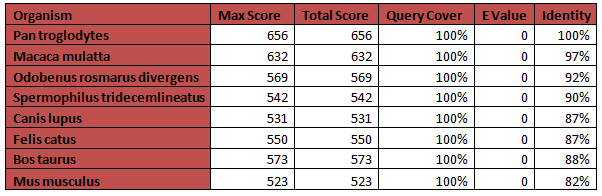
Using the Homo sapiens accession number found through Entrez, I was able to run a BLAST report finding the organisms with comparable NDN sequences to Homo sapiens. Using these homologs, I was able to conduct a Reciprocal BLAST so as to confirm my results. To the right is a table showcasing my BLAST results. The Max Score and Total Score comment on the quality of the sequence comparison, with higher being better (3). The Query Coverage allows us to know how much of the given sequence lines up with the Homo sapiens sequence it was compared to. The E Value shows statistical significance, with zero being the best option (3). Finally the Maximum Identity shows the percent identity between Homo sapiens and the given sequence based on amino acid-to-amino acid alignment (3).
Using the Homo sapiens accession number found through Entrez, I was able to run a BLAST report finding the organisms with comparable NDN sequences to Homo sapiens. Using these homologs, I was able to conduct a Reciprocal BLAST so as to confirm my results. To the right is a table showcasing my BLAST results. The Max Score and Total Score comment on the quality of the sequence comparison, with higher being better (3). The Query Coverage allows us to know how much of the given sequence lines up with the Homo sapiens sequence it was compared to. The E Value shows statistical significance, with zero being the best option (3). Finally the Maximum Identity shows the percent identity between Homo sapiens and the given sequence based on amino acid-to-amino acid alignment (3).

Macaca mulatta (Rhesus Macaque)
necdin Accession: NP_001165573 Length: 321 aa Odobenus rosmarus divergens (Walrus) necdin-like Accession: XP_004405540.1 Length: 325 aa Bos taurus (Domestic Cow) necdin Accession: NP_001014982.1 Length: 325 aa Spermophilus tridecemlineatus (Squirrel) necdin-like Accession: XP_005340621 Length: 325 |

Pan troglodytes (Common Chimpanzee)
necdin Accession: XP_510257 Length: 321 aa Canis lupus (Gray Wolf) necdin Accession: XP_005618297 Length: 325 aa Mus musculus (House Mouse) necdin Accession: NP_035012.2 Length: 325 aa Felis catus (Cat) necdin Accession: XP_011281123 Length: 327 |
Discussion of Necdin Homology
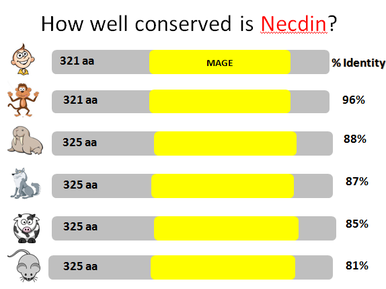
NDN is pretty well conserved which makes sense as it is a gene found only in mammals, and mammals have had a relatively recent common ancestor. Humans have a protein length of 321 amino acids along with the two other primates- rhesus macaques and chimpanzees. The other organisms, minus the cat which has a length of 377 amino acids, have a length of 325 amino acids. Because they are larger than humans, but still have a 100% Query Cover, we know that 321 of the amino acids are the same between, and these organisms have 4 extra amino acids, possibly leading to differences in functioning or localization.
NDN is pretty well conserved which makes sense as it is a gene found only in mammals, and mammals have had a relatively recent common ancestor. Humans have a protein length of 321 amino acids along with the two other primates- rhesus macaques and chimpanzees. The other organisms, minus the cat which has a length of 377 amino acids, have a length of 325 amino acids. Because they are larger than humans, but still have a 100% Query Cover, we know that 321 of the amino acids are the same between, and these organisms have 4 extra amino acids, possibly leading to differences in functioning or localization.
Phylogeny
What is Phylogeny?
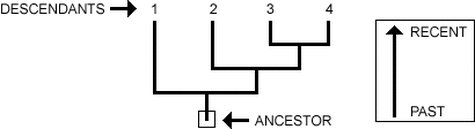
Phylogeny is the study of evolution and how different groups of organisms and species came to be. As we can see in the figure to the right, many organisms, labeled "descendants" can make up a phylogenetic tree. Each final line represents a separate species. Where each line intersects, a speciation event has occurred to a common ancestor preceding the two organisms. Those with closer intersections have had more recent common ancestors. Ultimately every organism can be traced to the same common ancestor. We can use phylogeny to track evolutionary relationships not only of organisms as a whole, but also the genetics of organisms, including the specific proteins. Below we will see an example of a phylogenetic tree sorted with the use of necdin protein sequences.
Phylogeny of Necdin
Using ClustalWOmega, it is possible to insert Fasta sequences of multiple species in order to align sequences and produce phylogenetic trees. With the aligned sequences, one can determine which amino acids are conserved throughout the protein and which changes may occur between species. This information is also applied to the phylogenetic trees which are determined by the amount of conservation. There are four types of phylogenetic tree layouts given by ClustalWOmega. Further information will be given below.
Information from ClustalWOmega looks like that shown below:
Phylogeny is the study of evolution and how different groups of organisms and species came to be. As we can see in the figure to the right, many organisms, labeled "descendants" can make up a phylogenetic tree. Each final line represents a separate species. Where each line intersects, a speciation event has occurred to a common ancestor preceding the two organisms. Those with closer intersections have had more recent common ancestors. Ultimately every organism can be traced to the same common ancestor. We can use phylogeny to track evolutionary relationships not only of organisms as a whole, but also the genetics of organisms, including the specific proteins. Below we will see an example of a phylogenetic tree sorted with the use of necdin protein sequences.
Phylogeny of Necdin
Using ClustalWOmega, it is possible to insert Fasta sequences of multiple species in order to align sequences and produce phylogenetic trees. With the aligned sequences, one can determine which amino acids are conserved throughout the protein and which changes may occur between species. This information is also applied to the phylogenetic trees which are determined by the amount of conservation. There are four types of phylogenetic tree layouts given by ClustalWOmega. Further information will be given below.
Information from ClustalWOmega looks like that shown below:
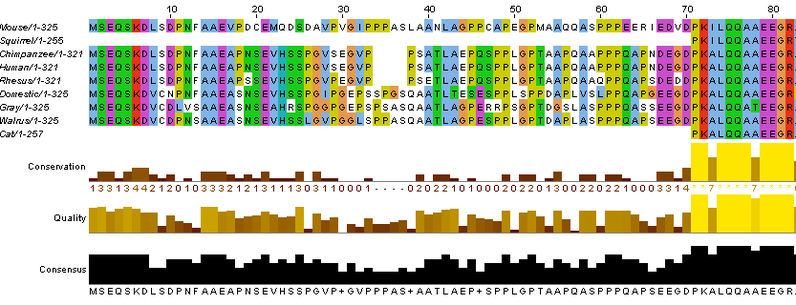
This is the beginning of the protein sequence of necdin compared across nine mammalian species. We can see that the beginning of the protein sequence is not very conserved between species, but further into the protein, it is quite conserved. Using this data, we can determine variants within the protein between species that may lead to variation in structure or function.
Average Distance Using Percent Identity
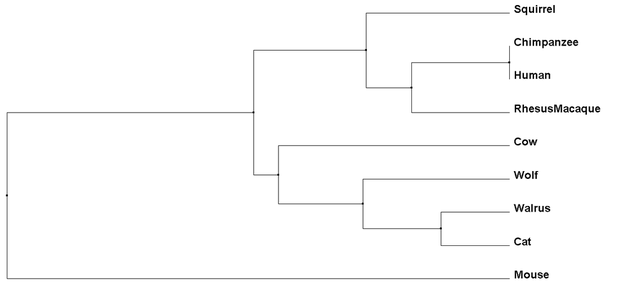
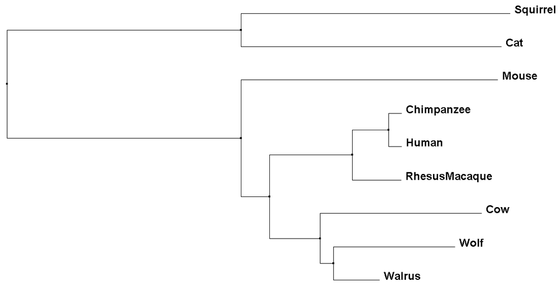
Percent Identity phylogenetic trees compare amino acid sequences of each species and determine which are the closest related to each other based on their conservation (4). Average Distance calculating then uses these determined relationships and gives all species the same length joining lines, showing the same amount of speciation for all organisms involved (4).
Percent Identity phylogenetic trees compare amino acid sequences of each species and determine which are the closest related to each other based on their conservation (4). Average Distance calculating then uses these determined relationships and gives all species the same length joining lines, showing the same amount of speciation for all organisms involved (4).
Average Distance Using BLOSUM62
BLOSUM62 is a scoring system that determines relatedness of species based on how well amino acid sequences are conserved and how likely this conservation would occur randomly (4). Like the tree above, the Average Distance calculating then uses these determined relationships and gives all species the same length joining lines, showing the same amount of speciation for all organisms involved (4).
BLOSUM62 is a scoring system that determines relatedness of species based on how well amino acid sequences are conserved and how likely this conservation would occur randomly (4). Like the tree above, the Average Distance calculating then uses these determined relationships and gives all species the same length joining lines, showing the same amount of speciation for all organisms involved (4).
Neighbor Joining Using Percent Identity
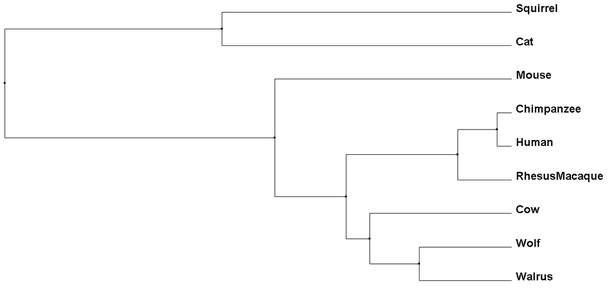
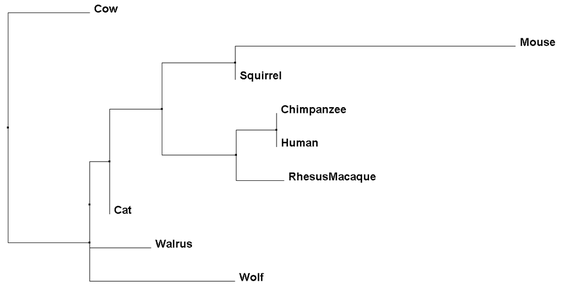
Percent Identity phylogenetic trees compare amino acid sequences of each species and determine which are the closest related to each other based on their conservation (4). Different from Average Distance trees, though, Neighbor Joining takes into account amount of change each species' protein sequence has had, leading to branches of different lengths (4).
Percent Identity phylogenetic trees compare amino acid sequences of each species and determine which are the closest related to each other based on their conservation (4). Different from Average Distance trees, though, Neighbor Joining takes into account amount of change each species' protein sequence has had, leading to branches of different lengths (4).
Neighbor Joining Using BLOSUM62
Again, BLOSUM62 is a scoring system that determines relatedness of species based on how well amino acid sequences are conserved and how likely this conservation would occur randomly (4). Different from Average Distance trees, though, Neighbor Joining takes into account amount of change each species' protein sequence has had, leading to branches of different lengths (4).
Again, BLOSUM62 is a scoring system that determines relatedness of species based on how well amino acid sequences are conserved and how likely this conservation would occur randomly (4). Different from Average Distance trees, though, Neighbor Joining takes into account amount of change each species' protein sequence has had, leading to branches of different lengths (4).
Discussion of Necdin Phylogeny
As we can see, each phylogenetic tree has a different total output, with some similarities seen throughout. For example, while the mouse is the outgroup for the Average Distance using Percent Identity Tree, it is more closely related to humans using the other three methods. Throughout all three methods, the Chimpanzee, Human, and Rhesus Macaque remain closely linked, as would be expected to their primate lineage. Finally, it is important to note that all of these organisms are mammals, as Necdin is only found in mammals. This limits amount of organisms for comparison of sequence and for further studies, but also denotes a relatively recent evolution of the protein.
As we can see, each phylogenetic tree has a different total output, with some similarities seen throughout. For example, while the mouse is the outgroup for the Average Distance using Percent Identity Tree, it is more closely related to humans using the other three methods. Throughout all three methods, the Chimpanzee, Human, and Rhesus Macaque remain closely linked, as would be expected to their primate lineage. Finally, it is important to note that all of these organisms are mammals, as Necdin is only found in mammals. This limits amount of organisms for comparison of sequence and for further studies, but also denotes a relatively recent evolution of the protein.
| fasta_sequences.docx | |
| File Size: | 12 kb |
| File Type: | docx |
Necdin MAGE Domain
What are Protein Domains?
A protein domain is a conserved and distinct portion of a protein that gives a particular structure or function, contributing to the end function of the protein (5). They may be conserved through many different types of proteins that may have very similar or very different functions (5). Some proteins will have many domains and others may have none or have just a single one. Domains can be used to sort proteins into different families with different functions as we will talk about with necdin's domain. Proteins may have domains that are not yet discovered, but still have great significance to the protein's overall structure.
What is Necdin's Domain?
Necdin has a single domain- the MAGE or the melanoma-associated antigen domain. This makes necdin a part of the MAGE superfamily of over 25 proteins in humans. The MAGE proteins are split into two types, Type I having expression in tumor cells and Type II having expression in differentiated cells, of which necdin is a part (6). MAGE domains are found in many organisms other than mammals including Drosophila and Aspergillus, but are absent in organisms such as C. elegans and unicellular organisms like yeast (7). The domain is typically 160-170 amino acids in length and is located on the necdin protein from amino acid 105 to 275 (6). It is well conserved among most mammals.
A protein domain is a conserved and distinct portion of a protein that gives a particular structure or function, contributing to the end function of the protein (5). They may be conserved through many different types of proteins that may have very similar or very different functions (5). Some proteins will have many domains and others may have none or have just a single one. Domains can be used to sort proteins into different families with different functions as we will talk about with necdin's domain. Proteins may have domains that are not yet discovered, but still have great significance to the protein's overall structure.
What is Necdin's Domain?
Necdin has a single domain- the MAGE or the melanoma-associated antigen domain. This makes necdin a part of the MAGE superfamily of over 25 proteins in humans. The MAGE proteins are split into two types, Type I having expression in tumor cells and Type II having expression in differentiated cells, of which necdin is a part (6). MAGE domains are found in many organisms other than mammals including Drosophila and Aspergillus, but are absent in organisms such as C. elegans and unicellular organisms like yeast (7). The domain is typically 160-170 amino acids in length and is located on the necdin protein from amino acid 105 to 275 (6). It is well conserved among most mammals.
Discussion of Necdin Domain
In necdin, the MAGE domain is required for binding to multiple other proteins, including p53, allowing its role in cell cycle regulation (6). Certain parts of the domain also play a part in nuclear matrix targetting and cell growth suppression (6). The MAGE takes up most of the human necdin protein and probably plays a big part in the main functioning of the protein. Also, because the MAGE is such a large domain, there is still much to be studied about it and its role both in necdin and also in the many different proteins it is a part of.
In necdin, the MAGE domain is required for binding to multiple other proteins, including p53, allowing its role in cell cycle regulation (6). Certain parts of the domain also play a part in nuclear matrix targetting and cell growth suppression (6). The MAGE takes up most of the human necdin protein and probably plays a big part in the main functioning of the protein. Also, because the MAGE is such a large domain, there is still much to be studied about it and its role both in necdin and also in the many different proteins it is a part of.
Necdin Interactions
What are Protein Interactions?
Proteins are the workhorses of the cell in that they carry out most of the functions of the cell. Some proteins are able to carry out functions by themselves, but others require the assistance of other proteins. When this occurs, it is considered to be a protein interaction. In studying a protein of interest, it can be very helpful to determine protein interactions so as to further identify possible functions of that protein based on known functions of those that it interacts with. STRING is a website that shows all known protein interactions based on published studies. The protein interaction networks are very dynamic due to constant publishing of studies.
Protein interactions are found using many different methods. Among these are co-immunoprecipitation & pull down, native gels, phage display, yeast two-hybrid systems, and tap-tagging (8). Tap tagging is a relatively new technique that uses high purification methods in order to be more accurate. This technique utilizes a tag to a protein of interest and measures it against other proteins to determine if they interact (9). It is especially effective, as it can identify complexes, and it does not localize to a particular point in the cell, as other methods do.
What are Necdin's Protein Interactions?
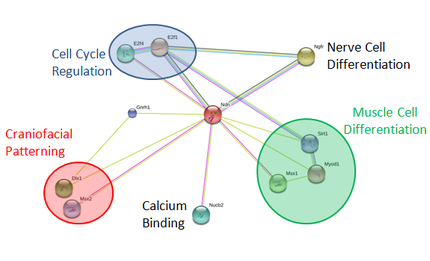
Using STRING, I was able to determine Necdin's already known protein interactions and separate them based on GO terms of each protein. Among these proteins, there are some that function in cell cycle regulation, nerve cell differentiation, craniofacial patterning, calcium binding, gonad differentiation, and muscle cell differentiation. In looking again into the STRING database, it was interesting to find that this interaction network has been updated in the times since taking this image. This means that necdin is an active protein of study, and there are sure to be more interactions, giving more understanding to the function of necdin.
Proteins are the workhorses of the cell in that they carry out most of the functions of the cell. Some proteins are able to carry out functions by themselves, but others require the assistance of other proteins. When this occurs, it is considered to be a protein interaction. In studying a protein of interest, it can be very helpful to determine protein interactions so as to further identify possible functions of that protein based on known functions of those that it interacts with. STRING is a website that shows all known protein interactions based on published studies. The protein interaction networks are very dynamic due to constant publishing of studies.
Protein interactions are found using many different methods. Among these are co-immunoprecipitation & pull down, native gels, phage display, yeast two-hybrid systems, and tap-tagging (8). Tap tagging is a relatively new technique that uses high purification methods in order to be more accurate. This technique utilizes a tag to a protein of interest and measures it against other proteins to determine if they interact (9). It is especially effective, as it can identify complexes, and it does not localize to a particular point in the cell, as other methods do.
What are Necdin's Protein Interactions?
Using STRING, I was able to determine Necdin's already known protein interactions and separate them based on GO terms of each protein. Among these proteins, there are some that function in cell cycle regulation, nerve cell differentiation, craniofacial patterning, calcium binding, gonad differentiation, and muscle cell differentiation. In looking again into the STRING database, it was interesting to find that this interaction network has been updated in the times since taking this image. This means that necdin is an active protein of study, and there are sure to be more interactions, giving more understanding to the function of necdin.
Discussion of Necdin Protein Interactions
Using this knowledge of Necdin protein interactions, we gain more insight into necdin's function within a cell. It should come as no surprise that necdin interacts with proteins functioning in cell cycle regulation and nerve cell differentiation, as these are two of its most studied roles. I found it's possible role in muscle cell differentiation to be rather interesting, due to PWS phenotypes of low muscle tone right from birth. This is an active area of study, and knowing these interactions could help in determining how necdin plays a role in this process. Also, the fact that it might play a role in the craniofacial phenotypes of PWS due to its interactions with these proteins is rather interesting, as I have not found any studies to expand on these interactions further.
Using this knowledge of Necdin protein interactions, we gain more insight into necdin's function within a cell. It should come as no surprise that necdin interacts with proteins functioning in cell cycle regulation and nerve cell differentiation, as these are two of its most studied roles. I found it's possible role in muscle cell differentiation to be rather interesting, due to PWS phenotypes of low muscle tone right from birth. This is an active area of study, and knowing these interactions could help in determining how necdin plays a role in this process. Also, the fact that it might play a role in the craniofacial phenotypes of PWS due to its interactions with these proteins is rather interesting, as I have not found any studies to expand on these interactions further.
Necdin Post-Translational Modifications
What are Post-Translational Modifications?
Post-translational modifications are chemical modifications to proteins that often change or add to the function of the protein (10). Examples of post-translational modifications are phosphorylation, glycosylation, ubiquitination, nitrosylation, methylation, and acetylation (10). In each of these processes, a functional group is added to the protein, allowing for a greater amount of protein complexity within the organism. These post-translational modifications can be found using many methods. One of these methods is mass spectrometry, which takes proteins and determines their sequence as well as any modifications based on weight. By gaining this information and studying further, we can better assess a protein's function and role in different processes.
What are Necdin's Post-Translational Modifications?
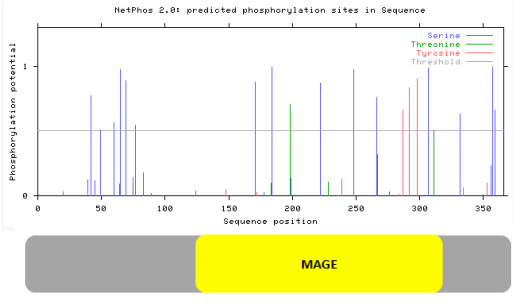
One post-translational modification that I focused on is phosphorylation because it is one of the most important and best studied post-translational modifications. Phosphorylation plays a role in regulating many different cell functions, including cell cycle and cell growth, both functions important for necdin (10). In order to find possible necdin phosphorylation sites, I used NetPhos 2.0, a server that predicts serine, threonine, and tyrosine phosphorylation sites for a protein.
Discussion of Necdin Post-Translational Modifications
In assessing necdin phosphorylation sites, I noticed that there are many within the MAGE domain, possibly contributing to the major roles of the MAGE domain. Also, the fact that there are so many possible phosphorylation sites shows that necdin could play a big role in regulation of other cell processes such as cell growth and cell cycle processes. We can look further at the phosphorylation sites by comparing the predicted sites of humans with the predicted sites of other organisms, such as mice to determine which are most well conserved. Those that are most conserved are probably important to necdin functioning and thus would provoke future studies. It could be beneficial to replace these phosphorylation site serines, threonines, or tyrosines with other amino acids to determine what the function of that phosphorylation site is.
Post-translational modifications are chemical modifications to proteins that often change or add to the function of the protein (10). Examples of post-translational modifications are phosphorylation, glycosylation, ubiquitination, nitrosylation, methylation, and acetylation (10). In each of these processes, a functional group is added to the protein, allowing for a greater amount of protein complexity within the organism. These post-translational modifications can be found using many methods. One of these methods is mass spectrometry, which takes proteins and determines their sequence as well as any modifications based on weight. By gaining this information and studying further, we can better assess a protein's function and role in different processes.
What are Necdin's Post-Translational Modifications?
One post-translational modification that I focused on is phosphorylation because it is one of the most important and best studied post-translational modifications. Phosphorylation plays a role in regulating many different cell functions, including cell cycle and cell growth, both functions important for necdin (10). In order to find possible necdin phosphorylation sites, I used NetPhos 2.0, a server that predicts serine, threonine, and tyrosine phosphorylation sites for a protein.
Discussion of Necdin Post-Translational Modifications
In assessing necdin phosphorylation sites, I noticed that there are many within the MAGE domain, possibly contributing to the major roles of the MAGE domain. Also, the fact that there are so many possible phosphorylation sites shows that necdin could play a big role in regulation of other cell processes such as cell growth and cell cycle processes. We can look further at the phosphorylation sites by comparing the predicted sites of humans with the predicted sites of other organisms, such as mice to determine which are most well conserved. Those that are most conserved are probably important to necdin functioning and thus would provoke future studies. It could be beneficial to replace these phosphorylation site serines, threonines, or tyrosines with other amino acids to determine what the function of that phosphorylation site is.
References:
1. http://www.britannica.com/EBchecked/topic/270557/homology
2. Wagner, 2007 http://www.nature.com.ezproxy.library.wisc.edu/nrg/journal/v8/n6/full/nrg2099.html
3. http://www.garlandscience.com/res/pdf/practicalbioinformatics_ch3.pdf
4. http://genetics564.weebly.com/homology--phylogeny.html
5. https://www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/protein-classification/what-are-protein-domains
6.http://onlinelibrary.wiley.com.ezproxy.library.wisc.edu/doi/10.1002/jcb.20345/full
7. http://onlinelibrary.wiley.com.ezproxy.library.wisc.edu/doi/10.1002/jnr.10160/full
8. https://www.lifetechnologies.com/us/en/home/life-science/protein-biology/protein-assays-analysis/protein-protein-interactions.html
9. http://www.ncbi.nlm.nih.gov/pubmed/11403571
10. https://www.lifetechnologies.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/overview-post-translational-modification.html
1. http://www.britannica.com/EBchecked/topic/270557/homology
2. Wagner, 2007 http://www.nature.com.ezproxy.library.wisc.edu/nrg/journal/v8/n6/full/nrg2099.html
3. http://www.garlandscience.com/res/pdf/practicalbioinformatics_ch3.pdf
4. http://genetics564.weebly.com/homology--phylogeny.html
5. https://www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/protein-classification/what-are-protein-domains
6.http://onlinelibrary.wiley.com.ezproxy.library.wisc.edu/doi/10.1002/jcb.20345/full
7. http://onlinelibrary.wiley.com.ezproxy.library.wisc.edu/doi/10.1002/jnr.10160/full
8. https://www.lifetechnologies.com/us/en/home/life-science/protein-biology/protein-assays-analysis/protein-protein-interactions.html
9. http://www.ncbi.nlm.nih.gov/pubmed/11403571
10. https://www.lifetechnologies.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/overview-post-translational-modification.html