This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison.
Model Organisms
What are Model Organisms?
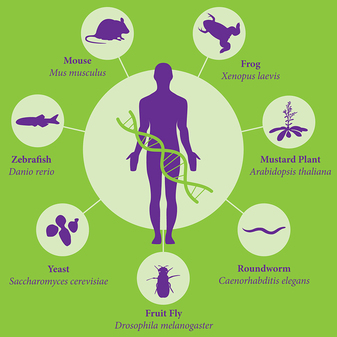
Model organisms are often used in research studies to give information about the human condition, especially in terms of diseases and drugs. Examples of model organisms are seen in the figure to the right. These include mice, frog, zebrafish, yeast, Drosophila, C. elegans, and the plant Arabidopsis. While doing studies on human development with zebrafish may seem silly due to vast differences in structure, many of the same development patterns in zebrafish are seen in humans as well. Indeed, as you may have seen in the Phylogeny section of this website, all organisms came from the same common ancestor making many processes conserved between species and allowing us to use these models to gain further information without using humans. This is especially helpful in genetic studies, as we can manipulate the genome in specific ways in different model organisms so as to find the phenotype and genotype of different genetic disorders. Many techniques can be used to manipulate these organisms, and many more continue to be discovered as scientists look further into the specificities of different systems.
Model organisms are often used in research studies to give information about the human condition, especially in terms of diseases and drugs. Examples of model organisms are seen in the figure to the right. These include mice, frog, zebrafish, yeast, Drosophila, C. elegans, and the plant Arabidopsis. While doing studies on human development with zebrafish may seem silly due to vast differences in structure, many of the same development patterns in zebrafish are seen in humans as well. Indeed, as you may have seen in the Phylogeny section of this website, all organisms came from the same common ancestor making many processes conserved between species and allowing us to use these models to gain further information without using humans. This is especially helpful in genetic studies, as we can manipulate the genome in specific ways in different model organisms so as to find the phenotype and genotype of different genetic disorders. Many techniques can be used to manipulate these organisms, and many more continue to be discovered as scientists look further into the specificities of different systems.
CRISPR
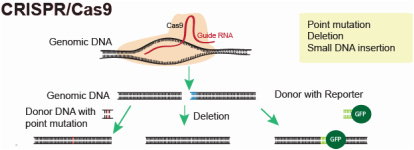
One of these techniques is called CRISPR-Cas9. This technique started being used in 2010, soon after it was found being used by microbes as an immune system adaptation to get rid of unwanted viral DNA within their genome. This system can be used to effectively introduce deletions or insertions into the genetic code in order to create knockout or knock-in organisms. Simply put, a guide RNA attached to Cas9 targets the sequence of choice (3). The Cas9 then creates a double-stranded break at this point, allowing the bases at this point to be taken out and/or new bases to be added (3). It is possible that this technique may become the most used way to create knockout mutations in organisms, replacing homologous recombination and RNAi, among others as it is a very effective way to completely knockout the gene in question.
Unfortunately a CRISPR system has not yet been made for NDN, but it is possible that the future may bring such a product in order to do further studies into the effects of NDN deletion in creating PWS symptoms.
PWS/NDN Model Organisms
Of the main model organisms in use today, NDN is only found in the mammals- mice and rats. Most studies of NDN expression and lack of expression in PWS has been studied in mice thus far. These mice have been used to determine normal NDN expression and role in the organism, as well as what deletion of NDN will cause. I looked up NDN model mice on the Mouse Genome Informatics Website and found that there are at least six posted targetted NDN deletion models used currently. NDN-targetted knockout mice generally have partial neonatal lethality, respiratory distress, abnormal behavior, and abnormal nervous system morphology and physiology (1). However, each of these mutated mice is done on a different genetic background, leading to different phenotypic outcomes including a strain that showed no symptoms of Prader-Willi Syndrome. This is because, like humans, mice of different genetic backgrounds can react to the same mutation differently (2). Because of this, having different disease models is important to determining the full extent of how a protein or a mutation works within a disease.
Discussion of PWS/NDN Model Organisms
Mice have been a great source of information about Prader-Willi Syndrome thus far. Many different techniques have already been used to gather information about normal NDN expression as well as NDN deletions. Because we are using mice as a model organism, there are some limitations including the fact that necdin is expressed in different tissues in humans and mice, for while mice have expression solely in the brain, humans have expression almost ubiquitously (4). This leads to many areas of future study in order to completely discern what the true function of necdin is in creating PWS symptoms. While there is much already known about NDN function, there is still a long way to go!
Look below for a short video explaining why we should trust studies done on model organisms like mice!
One of these techniques is called CRISPR-Cas9. This technique started being used in 2010, soon after it was found being used by microbes as an immune system adaptation to get rid of unwanted viral DNA within their genome. This system can be used to effectively introduce deletions or insertions into the genetic code in order to create knockout or knock-in organisms. Simply put, a guide RNA attached to Cas9 targets the sequence of choice (3). The Cas9 then creates a double-stranded break at this point, allowing the bases at this point to be taken out and/or new bases to be added (3). It is possible that this technique may become the most used way to create knockout mutations in organisms, replacing homologous recombination and RNAi, among others as it is a very effective way to completely knockout the gene in question.
Unfortunately a CRISPR system has not yet been made for NDN, but it is possible that the future may bring such a product in order to do further studies into the effects of NDN deletion in creating PWS symptoms.
PWS/NDN Model Organisms
Of the main model organisms in use today, NDN is only found in the mammals- mice and rats. Most studies of NDN expression and lack of expression in PWS has been studied in mice thus far. These mice have been used to determine normal NDN expression and role in the organism, as well as what deletion of NDN will cause. I looked up NDN model mice on the Mouse Genome Informatics Website and found that there are at least six posted targetted NDN deletion models used currently. NDN-targetted knockout mice generally have partial neonatal lethality, respiratory distress, abnormal behavior, and abnormal nervous system morphology and physiology (1). However, each of these mutated mice is done on a different genetic background, leading to different phenotypic outcomes including a strain that showed no symptoms of Prader-Willi Syndrome. This is because, like humans, mice of different genetic backgrounds can react to the same mutation differently (2). Because of this, having different disease models is important to determining the full extent of how a protein or a mutation works within a disease.
Discussion of PWS/NDN Model Organisms
Mice have been a great source of information about Prader-Willi Syndrome thus far. Many different techniques have already been used to gather information about normal NDN expression as well as NDN deletions. Because we are using mice as a model organism, there are some limitations including the fact that necdin is expressed in different tissues in humans and mice, for while mice have expression solely in the brain, humans have expression almost ubiquitously (4). This leads to many areas of future study in order to completely discern what the true function of necdin is in creating PWS symptoms. While there is much already known about NDN function, there is still a long way to go!
Look below for a short video explaining why we should trust studies done on model organisms like mice!
Gene Expression
What is Gene Expression?
DNA encodes mRNA which encodes proteins which go into the cell to carry out a function of some sort. The amount of protein present often determines how and when a function is carried out. When a function is being carried out, this is called gene expression. Indeed, just because a person has a gene inside of all their cells, it does not mean that all of these genes are being expressed all of the time. There are many mechanisms which regulate whether or not a gene is being expressed in a cell at any given time. Gene expression often occurs at the protein level, but often it is easier to measure it at the mRNA or even DNA level. There are many methods to measuring gene expression, including Northern Blots, Western Blots, ELISA, and RT-qPCR. One that we are going to focus on is the microarray.
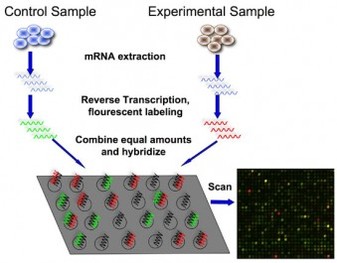
The microarray is an effective technique in measuring mRNA levels within a sample. Researchers will set up a microarray with known sequences of DNA and then run the unknown mRNA sequences through. The mRNA sequences will bind to DNA to which it is complementary, and through the viewing technology of the microarray, it is possible to determine the quantity of mRNA bound to the known DNA sequence. This technique can be applied to many types of experiments, including those looking to determine proteins involved in specific diseases and those working towards discovery of new drugs or genes (5).
DNA encodes mRNA which encodes proteins which go into the cell to carry out a function of some sort. The amount of protein present often determines how and when a function is carried out. When a function is being carried out, this is called gene expression. Indeed, just because a person has a gene inside of all their cells, it does not mean that all of these genes are being expressed all of the time. There are many mechanisms which regulate whether or not a gene is being expressed in a cell at any given time. Gene expression often occurs at the protein level, but often it is easier to measure it at the mRNA or even DNA level. There are many methods to measuring gene expression, including Northern Blots, Western Blots, ELISA, and RT-qPCR. One that we are going to focus on is the microarray.
The microarray is an effective technique in measuring mRNA levels within a sample. Researchers will set up a microarray with known sequences of DNA and then run the unknown mRNA sequences through. The mRNA sequences will bind to DNA to which it is complementary, and through the viewing technology of the microarray, it is possible to determine the quantity of mRNA bound to the known DNA sequence. This technique can be applied to many types of experiments, including those looking to determine proteins involved in specific diseases and those working towards discovery of new drugs or genes (5).
NDN Gene Expression
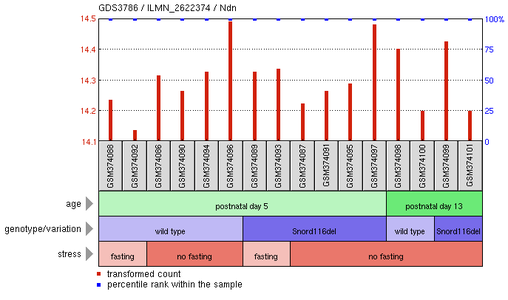
It is possible to search Gene Expression Omnibus, or GEO, for past studies that have searched for expression of a certain gene within different conditions. In searching for NDN and Prader-Willi Syndrome, I found a study looking at the Snord116del mutant hypothalamus response to neonatal starvation. This study looked at different protein levels during fasting and non-fasting times either in the controlled wildtype or in the Snord116 mutant, another mutation found connected to Prader-Willi Syndrome. Individual results of this study are seen to the right. Each gray box denotes a specific group of mice in that condition. The conditions of the study are seen below these gray boxes and the amounts of NDN present for each sample at these different conditions are represented by the red lines stemming from each gray box.
It is possible to search Gene Expression Omnibus, or GEO, for past studies that have searched for expression of a certain gene within different conditions. In searching for NDN and Prader-Willi Syndrome, I found a study looking at the Snord116del mutant hypothalamus response to neonatal starvation. This study looked at different protein levels during fasting and non-fasting times either in the controlled wildtype or in the Snord116 mutant, another mutation found connected to Prader-Willi Syndrome. Individual results of this study are seen to the right. Each gray box denotes a specific group of mice in that condition. The conditions of the study are seen below these gray boxes and the amounts of NDN present for each sample at these different conditions are represented by the red lines stemming from each gray box.
Discussion of NDN Gene Expression
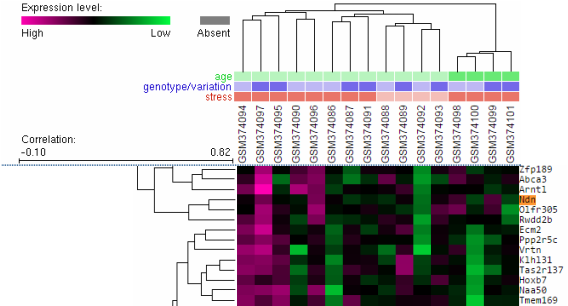
An example of what a microarray looks like when finished is seen on the left. This is a small portion of the entire microarray done for the study talked about above. The conditions above the array are the same as those below the samples above. We can see that there is a measured amount of expression depending on the different conditions. To the right, NDN is highlighted in orange. Between this colored expression and the figure above, we can see that NDN usually increases with no fasting in the wildtype (normal conditions), but there is some variablitiy. With the Snord116 deletion, NDN levels are similar to the wiltype.
While many other types of gene expression studies have been done in regards to necdin and PWS, this was the only one recorded in GEO. Looking toward the future, as more studies are done, it is possible that there will be a microarray studying necdin deletion and how other proteins are affected.
An example of what a microarray looks like when finished is seen on the left. This is a small portion of the entire microarray done for the study talked about above. The conditions above the array are the same as those below the samples above. We can see that there is a measured amount of expression depending on the different conditions. To the right, NDN is highlighted in orange. Between this colored expression and the figure above, we can see that NDN usually increases with no fasting in the wildtype (normal conditions), but there is some variablitiy. With the Snord116 deletion, NDN levels are similar to the wiltype.
While many other types of gene expression studies have been done in regards to necdin and PWS, this was the only one recorded in GEO. Looking toward the future, as more studies are done, it is possible that there will be a microarray studying necdin deletion and how other proteins are affected.
Chemical Genetics

What is Chemical Genetics?
Chemical Genetics is a technique that uses small molecules to discover biological processes of different proteins (6). The setup can be very similar to the microarray seen above, except instead of known DNA sequences, you set the microarray up with known small molecules. As proteins are run through the array, various ones will be able to bind to various small molecules. By using this information, researchers can then look further into the relationship between the small molecule and the protein that has bound, determining biological pathways or even possible drugs (6). Many small molecules have already been cataloged in libraries for use of researchers along with known interaction partners. Using the knowledge of small molecules, how they work, and what they interact with, it is possible to determine functions for vast amounts of proteins
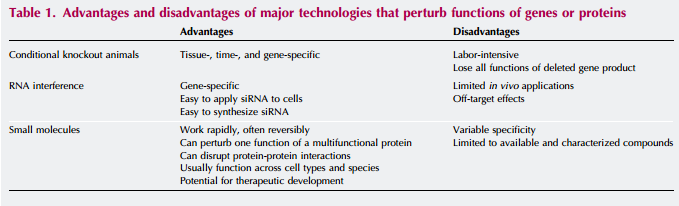
Also, small molecules can be added to the list of genetic techniques used to manipulate organisms in vivo. Small molecules can be inserted into organisms to perturb the system, creating a quick and reversible study of how the system works with specific proteins out of the mix. Below is a quick look at a couple of these other techniques in comparison to small molecule use.
NDN Chemical Genetics
It is possible to find small molecule/protein interactions already discovered using PubChem. If a person already knows possible drugs or small molecules which interact with the protein, they can search by that, or a person can search for the specific protein or group of proteins that they are studying. Also, it is possible to look at specific diseases and whether there have been any small molecules aligned with them by using the Bioassays tab. Unfortunately, in looking for NDN and/or Prader-Willi Syndrome, I was not able to find any small molecules that have been discovered to this point.
Discussion of NDN Chemical Genetics
A lack of information here does not denote that there are no small molecules that do bind to necdin. It is quite possible that there are different types of small molecules that do bind, especially as necdin is typically involved with binding other proteins or the DNA directly. As further studies occur, a chemical genetics assay could be used to determine further pathways of NDN and, possibly, drugs that could be used as treatment for specific symptoms of Prader-Willi Syndrome.
Chemical Genetics is a technique that uses small molecules to discover biological processes of different proteins (6). The setup can be very similar to the microarray seen above, except instead of known DNA sequences, you set the microarray up with known small molecules. As proteins are run through the array, various ones will be able to bind to various small molecules. By using this information, researchers can then look further into the relationship between the small molecule and the protein that has bound, determining biological pathways or even possible drugs (6). Many small molecules have already been cataloged in libraries for use of researchers along with known interaction partners. Using the knowledge of small molecules, how they work, and what they interact with, it is possible to determine functions for vast amounts of proteins
Also, small molecules can be added to the list of genetic techniques used to manipulate organisms in vivo. Small molecules can be inserted into organisms to perturb the system, creating a quick and reversible study of how the system works with specific proteins out of the mix. Below is a quick look at a couple of these other techniques in comparison to small molecule use.
NDN Chemical Genetics
It is possible to find small molecule/protein interactions already discovered using PubChem. If a person already knows possible drugs or small molecules which interact with the protein, they can search by that, or a person can search for the specific protein or group of proteins that they are studying. Also, it is possible to look at specific diseases and whether there have been any small molecules aligned with them by using the Bioassays tab. Unfortunately, in looking for NDN and/or Prader-Willi Syndrome, I was not able to find any small molecules that have been discovered to this point.
Discussion of NDN Chemical Genetics
A lack of information here does not denote that there are no small molecules that do bind to necdin. It is quite possible that there are different types of small molecules that do bind, especially as necdin is typically involved with binding other proteins or the DNA directly. As further studies occur, a chemical genetics assay could be used to determine further pathways of NDN and, possibly, drugs that could be used as treatment for specific symptoms of Prader-Willi Syndrome.
References:
1)http://www.informatics.jax.org/marker/MGI:97290
2) http://research.jax.org/mousegenetics/advantages/genetic-background-effects.html
3) https://www.addgene.org/CRISPR/guide/
4) Macdonald, H., & Wevrick, R. (1997). The Necdin Gene is Deleted in Prader-Willi Syndrome and is Imprinted in Human and Mouse. Human Molecular Genetics,6(11), 1873-1878.
5) http://www.premierbiosoft.com/tech_notes/microarray.html
6) http://www.pnlab.org/publications/documents/5700853a.pdf
1)http://www.informatics.jax.org/marker/MGI:97290
2) http://research.jax.org/mousegenetics/advantages/genetic-background-effects.html
3) https://www.addgene.org/CRISPR/guide/
4) Macdonald, H., & Wevrick, R. (1997). The Necdin Gene is Deleted in Prader-Willi Syndrome and is Imprinted in Human and Mouse. Human Molecular Genetics,6(11), 1873-1878.
5) http://www.premierbiosoft.com/tech_notes/microarray.html
6) http://www.pnlab.org/publications/documents/5700853a.pdf
